The Wistar Institute’s Dr. Noam Auslander was awarded a $600,000 Women Scientists Innovation Award for Cancer Research grant from the V Foundation for Cancer Research to support the next three years of her research, which will use proteins of gut bacteria to predict immunotherapy benefit in melanoma.
Tag: Computational Biology
LJI scientists develop new method to match genes to their molecular ‘switches’
LA JOLLA, CA—Scientists at La Jolla Institute for Immunology (LJI) have developed a new computational method for linking molecular marks on our DNA to gene activity. Their work may help researchers connect genes to the molecular “switches” that turn them on or off. This research, published in Genome Biology, is an important step toward harnessing machine learning approaches to better understand links between gene expression and disease development.
Scientists pioneer autonomous robotic method for studying liquids suspended in air
Researchers at the U.S. Department of Energy’s Argonne National Laboratory have used robots and artificial intelligence to dramatically speed up data collection and analysis in X-ray studies of liquids.
Scientists find core regulatory circuit controlling identity of aggressive leukemia
Scientists at St. Jude Children’s Research Hospital and Dana-Farber Cancer Institute found a small set of proteins that maintain anaplastic large cell lymphoma (ALCL) identity, representing potential future therapeutic targets.
Researchers from IRB Barcelona and Sant Joan de Déu unveil the origin of second pediatric cancers and chemotherapy-induced mutations in healthy tissues
A team from IRB Barcelona and Sant Joan de Déu Hospital · IRSJD has studied the cases of four children who have experienced two cancers during childhood and has addressed the origin of the second tumour.
The Wistar Institute Recruits Dr. Avi Srivastava as Assistant Professor
Wistar is pleased to announce the recruitment of Avi Srivastava, Ph.D., to the Ellen and Ronald Caplan Cancer Center, where he joins Wistar’s Gene Expression and Regulation Program as an Assistant Professor.
Department of Energy Announces $4.7 Million for Research on Integrative Computational Tools for Systems Biology Research
Today, the U.S. Department of Energy (DOE) announced $4.7 million in funding for five new research projects in computational biology. These projects will develop new software and analytical tools to manage the growing quantities of genomics and other data stemming from the study of microbes and other biological systems.

Jinghui Zhang, PhD, elected Fellow of the International Society for Computational Biology
Former Chair of Computational Biology honored with induction into Class of 2023 Fellows.
Researchers use Argonne X-rays to find the best antibodies
Antibody therapies are only effective if the antibodies do what we want them to do. This research can help scientists determine if an antibody is likely to stick to something other than the intended target, which should lessen the amount of time wasted with overly sticky antibodies.
2023 Warren Alpert Foundation Prize Honors Pioneer in Computational Biology
The 2023 Warren Alpert Foundation Prize has been awarded to scientist David J. Lipman for his visionary work in the conception, design, and implementation of computational tools, databases, and infrastructure that transformed the way biological information is analyzed and accessed freely and rapidly around the world.
New Computational Tool Identifies Novel Targets for Cancer Immunotherapy
Researchers at Children’s Hospital of Philadelphia (CHOP) and the University of California, Los Angeles (UCLA) have developed a computational platform capable of discovering tumor antigens derived from alternative RNA splicing, expanding the pool of cancer immunotherapy targets. The tool, called “Isoform peptides from RNA splicing for Immunotherapy target Screening” (IRIS), was described in a paper published today in the Proceedings of the National Academy of Sciences.

Computational biology superstar Sanju Sinha joins Sanford Burnham Prebys
Sanju Sinha, Ph.D., has joined Sanford Burnham Prebys as an assistant professor in the Cancer Molecular Therapeutics Program to continue his research on cancer development and drug discovery. He comes to Sanford Burnham Prebys from the Cancer Data Science Lab at the National Cancer Institute (NCI), where he trained to leverage the power of artificial intelligence to discover new drugs to prevent cancer.
New algorithm can predict diabetic kidney disease
Researchers from Sanford Burnham Prebys and the Chinese University of Hong Kong have developed a computational approach to predict whether a person with type 2 diabetes will develop kidney disease. The findings could help doctors prevent or better manage kidney disease in people with type 2 diabetes.
New $17 million grant establishes LJI as global hub for immunology data curation and analysis
A new grant of over $17 million from the National Institute of Allergy and Infectious Diseases (NIAID) has established La Jolla Institute for Immunology (LJI) as the leading institute for human immunology data curation, analysis, and dissemination. With this funding, LJI has taken the helm of the Human Immunology Project Consortium Data Coordinating Center, a critical tool in the effort to fuel scientific collaboration in immunoprofiling and highlight findings from the overall Human Immunology Project Consortium (HIPC).
Q&A with Madhurima Vardhan, Argonne’s Margaret Butler Fellow
As the latest recipient of the Argonne Leadership Computing Facility’s Margaret Butler Fellowship, Madhurima Vardhan will use Argonne’s supercomputing and AI to advance biomedical research.
Getting to the Heart of Chemotherapeutic Cardiotoxicity
On any given Tuesday, you will find Brian C. Jensen, MD, cardiologist and physician-scientist, tending to patients in his cardio-oncology clinic. His schedule is packed to the brim with cancer patients. But not patients with heart cancer. The largest number of patients he sees are cancer patients who have developed, or are at risk of developing, heart damage in response to their chemotherapy regimens.
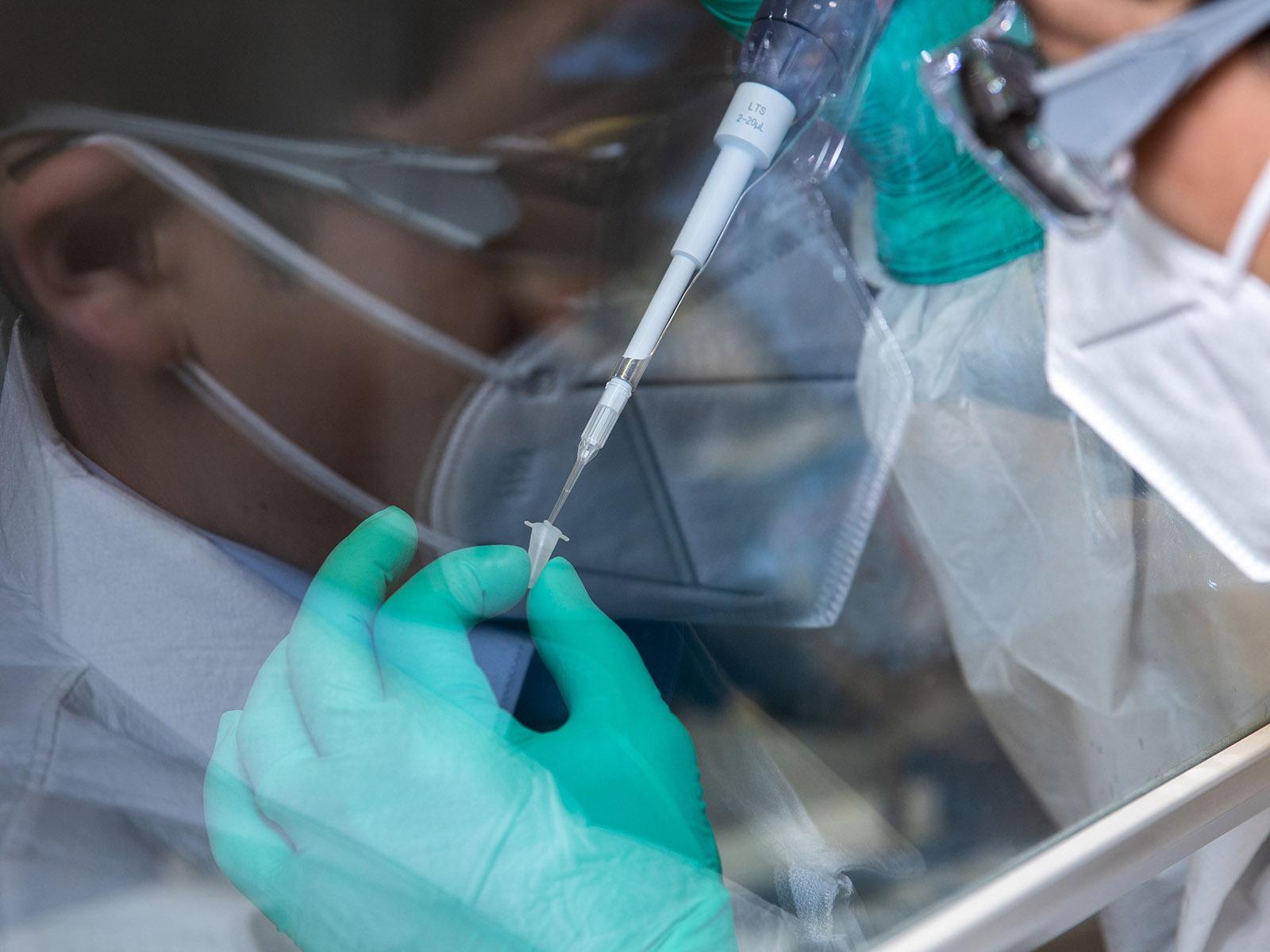
Big Data Analysis Powers the Fight Against Alzheimer’s
New research findings published in Science Advances (November 2022), help explain the progression of Alzheimer-related dementia in each patient. The findings outline a biological classification system that predicts disease severity.
Adapting language models to track virus variants
Groundbreaking research by Argonne National Laboratory finds new method to quickly identify COVID-19 virus variants. Their work wins the Gordon Bell Special Prize.
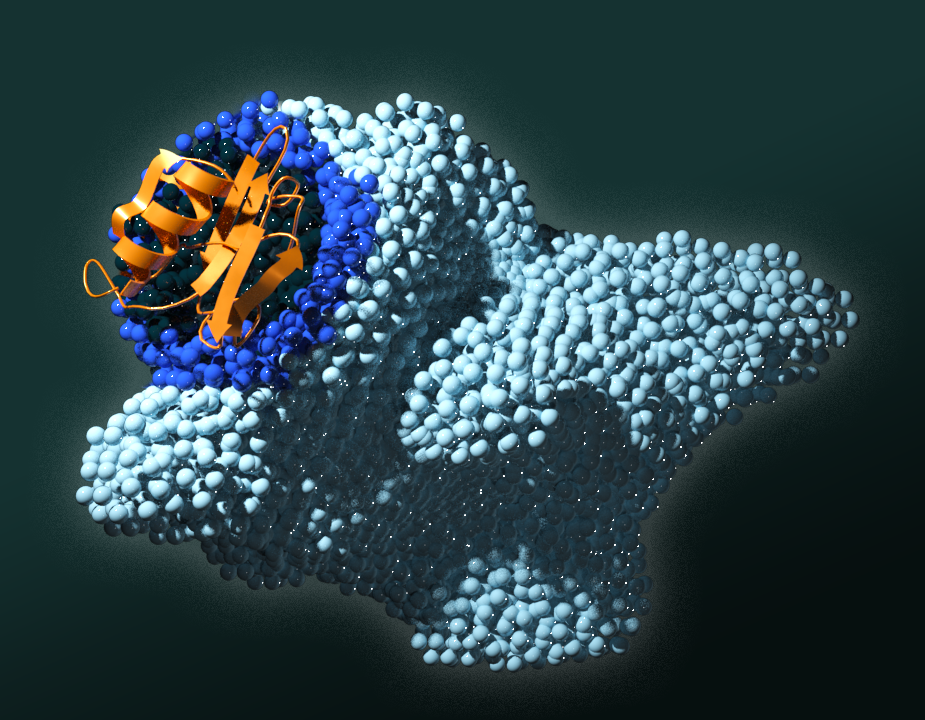
Scientists discover cancer trigger that could spur targeted drug therapies
Researchers at the Department of Energy’s Oak Ridge National Laboratory have definitively linked the function of a specific domain of proteins important in plant-microbe biology to a cancer trigger in humans, knowledge that had eluded scientists for decades.
UCSF Awarded $67.5 Million to Develop New Antiviral Therapies
Scientists at the UC San Francisco (UCSF) Quantitative Biosciences Institute (QBI) and the QBI Coronavirus Research Group (QCRG) have been awarded $67.5 million from the National Institute of Allergy and Infectious Diseases (NIAID) to support its mission of pandemic preparedness.
CHOP Researchers Develop New Computational Tool to Interpret Clinical Significance of Cancer Mutations
Researchers at Children’s Hospital of Philadelphia (CHOP) have developed a new tool to help researchers interpret the clinical significance of somatic mutations in cancer. The tool, known as CancerVar, incorporates machine learning frameworks to go beyond merely identifying somatic cancer mutations and interpret the potential significance of those mutations in terms of cancer diagnosis, prognosis, and targetability. A paper describing CancerVar was published today in Science Advances.
Sloan Kettering Institute’s Dana Pe’er Named Howard Hughes Medical Institute Investigator
Dana Pe’er, PhD, computational biologist and lab head at Memorial Sloan Kettering Cancer Center’s (MSK) Sloan Kettering Institute (SKI), is one of 33 biomedical researchers named a Howard Hughes Medical Institute (HHMI) investigator today.
Argonne collaborations bring computational tools to the forefront of COVID-19 research
Argonne, industry and academia collaborate to bring innovative AI and simulation tools to the COVID-19 battlefront.
AI Predicts How Patients with Viral Infections, Including COVID-19, Will Fare
UC San Diego School of Medicine researchers discovered gene expression patterns associated with pandemic viral infections, providing a map to help define patients’ immune responses, measure disease severity, predict outcomes and test therapies — for current and future pandemics.
DOE scientists deploy creativity, speed to disrupt COVID-19
An ORNL-led team comprising researchers from multiple DOE national laboratories is using artificial intelligence and computational screening techniques – in combination with experimental validation – to identify and design five promising drug therapy approaches to target the SARS-CoV-2 virus.
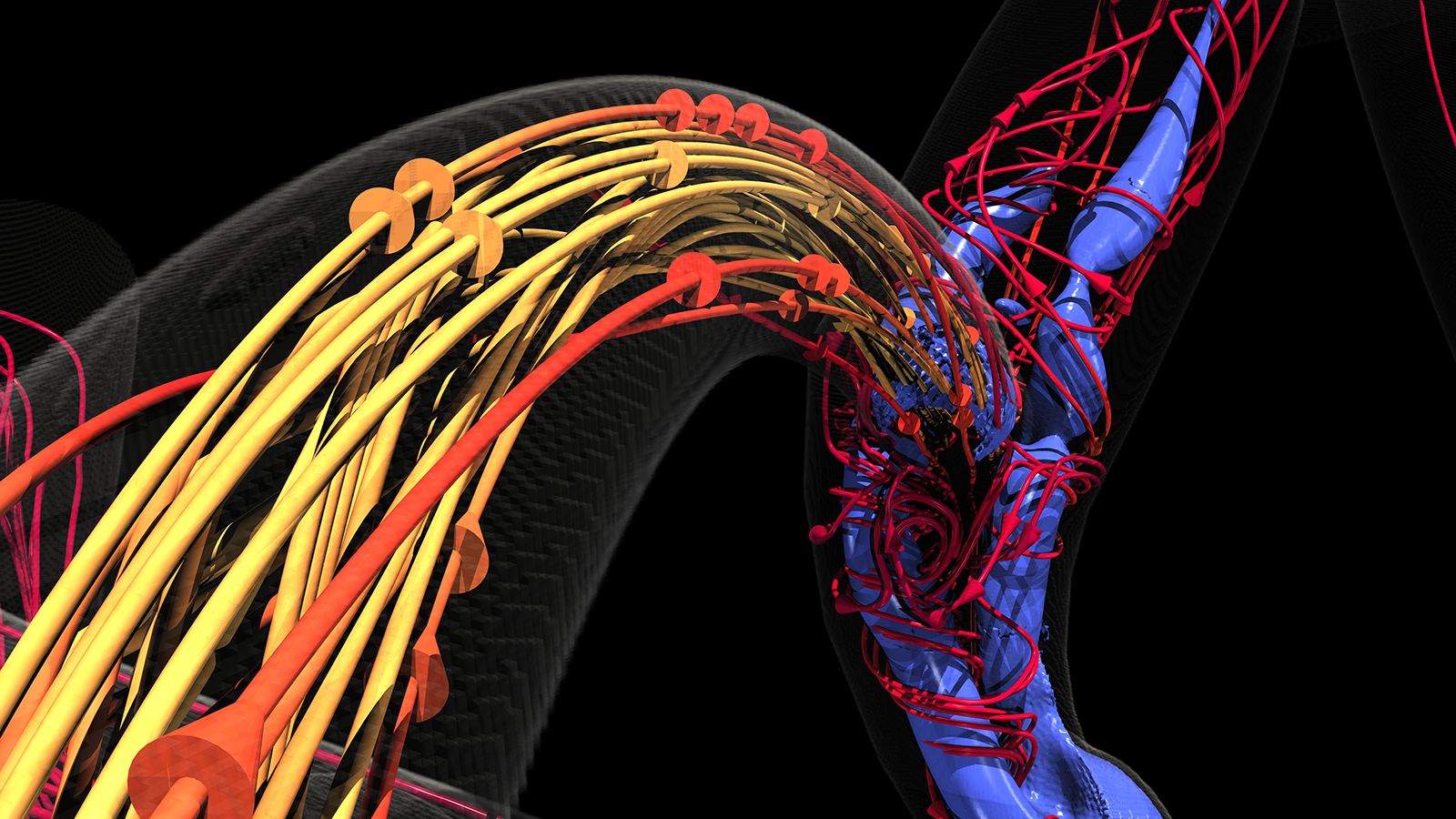
Preparing for exascale: Aurora supercomputer to help scientists visualize the spread of cancer
In advance of Argonne’s Aurora exascale supercomputer, Duke University assistant professor Amanda Randles is leading a new study to analyze cancer metastasis using HARVEY, a code that simulates blood vessels within the human body.
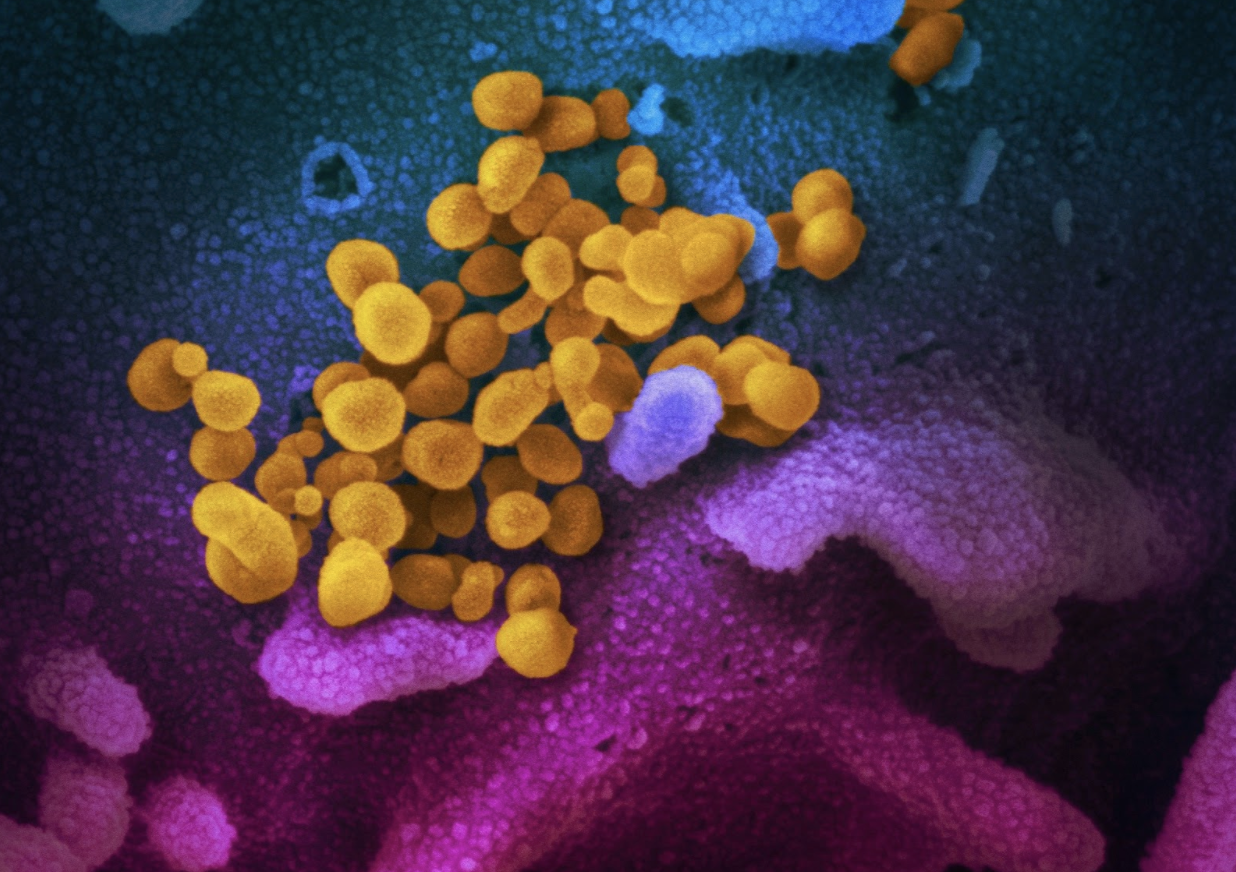
Science Snapshots: COVID-19, power outages, Alzheimer’s disease, and optical antennas
March Science Snapshots from Berkeley Lab
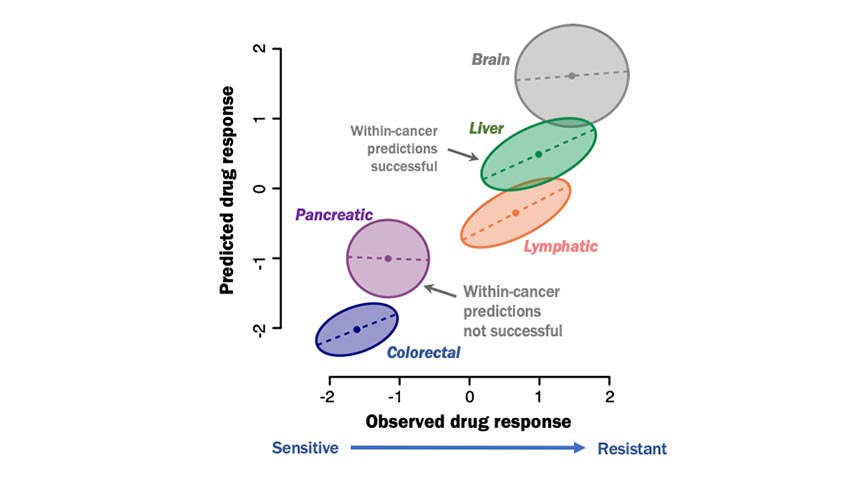
New Model Predicts Cancer Drug Efficacy Across and Within Cancer Types
As large multi-cancer datasets become more important for predicting who may benefit from cancer drugs, a new model better accounts for potentially overlooked variation.

The AI-driven initiative that’s hastening the discovery of drugs to treat COVID-19
Ten organizations have created a pipeline of artificial intelligence and simulation tools to narrow the search for drug candidates that can inhibit SARS-CoV-2.
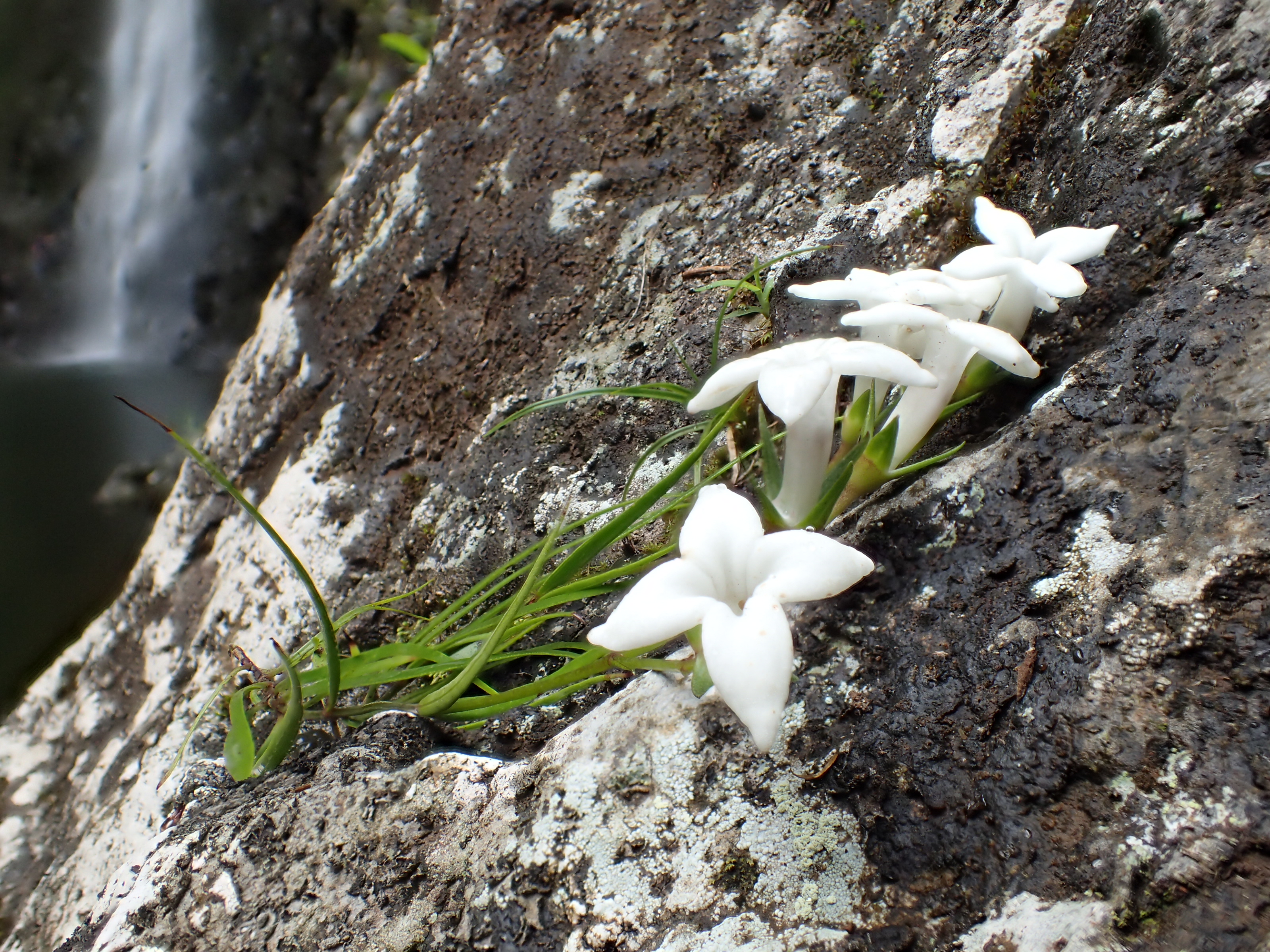

Biologists awarded NSF grant to model evolution of Hawaiian plants
The research team will focus on 12 understudied plant groups — including Kadua, Wikstroemia and Psychotria — using new sequencing and modeling techniques to gain a broader idea of how Hawaiian plant diversity originated. The project includes conservation and educational components as well, including collecting new wild specimens for the National Tropical Botanical Garden herbarium.
10 ways Argonne science is combatting COVID-19
Argonne scientists and research facilities have made a difference in the fight against COVID-19 in the year since the first gene sequence for the virus was published.
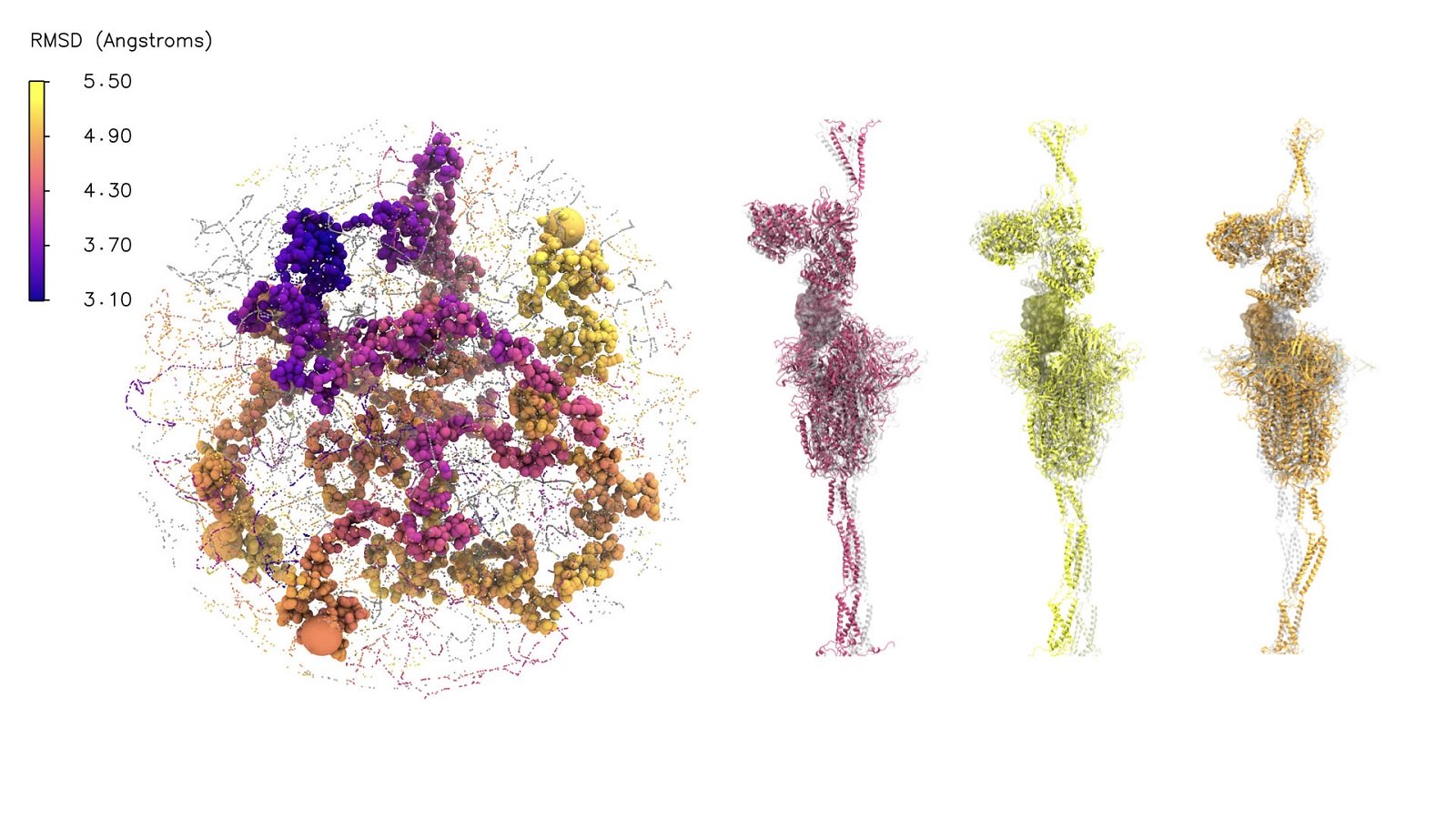
Argonne AI methods unravel mysteries of SARS-CoV-2 viral-human cell interaction
Using a combination of AI and supercomputing resources, Argonne researchers are examining the dynamics of the SARS-CoV-2 spike protein to determine how it fuses with the human host cell, advancing the search for drug treatments.

Uncovering Novel Genomes from Earth’s Microbiomes
Reported in Nature Biotechnology, the known diversity of bacteria and archaea has been expanded by 44% through a publicly available collection of more than 52,000 microbial genomes from environmental samples resulting from a JGI-led collaboration involving more than 200 scientists around the world.
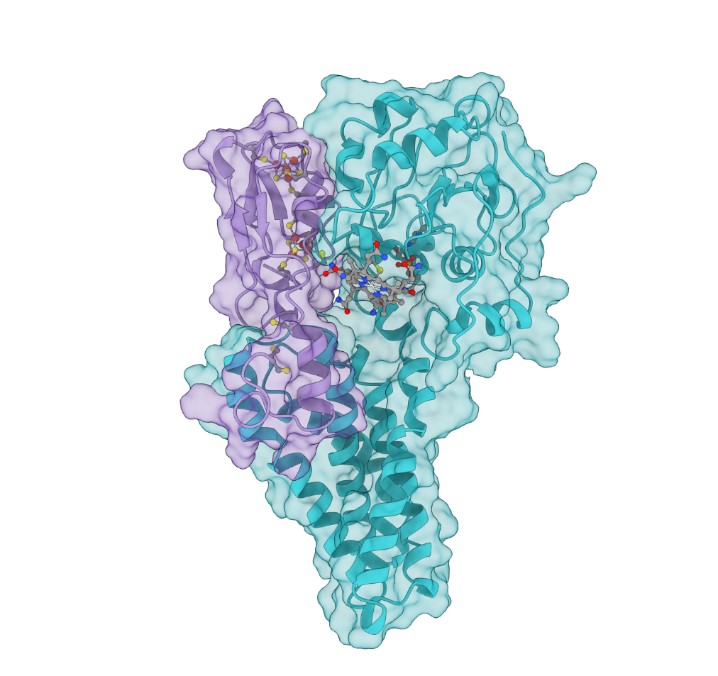
Scientists create protein models to explore toxic methylmercury formation
A team led by the Department of Energy’s Oak Ridge National Laboratory created a computational model of the proteins responsible for the transformation of mercury to toxic methylmercury, marking a step forward in understanding how the reaction occurs and how mercury cycles through the environment.
New Computational Model by CHOP Researchers Identifies Noncoding Mutations Across Five Pediatric Cancers
Researchers at Children’s Hospital of Philadelphia (CHOP) have developed a new computational algorithm that has, for the first time, identified a spectrum of mutations in the noncoding portion of the human genome across five major pediatric cancers. The study, which was published today in Science Advances, used the algorithm to analyze more than 500 pediatric cancer patients’ mutations and gene expression profiles to develop a comprehensive list of potentially cancer-causing mutations.
DOE Awards $5 Million for Computational Biology Research
The U.S. Department of Energy (DOE) announced $5 million in funding for six new research projects in computational biology.
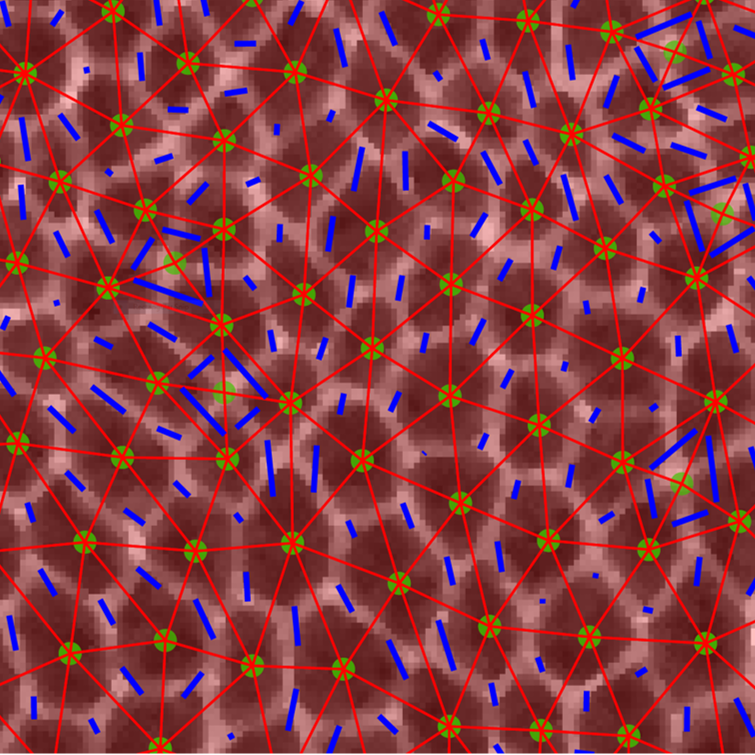
New View on How Tissues Flow in the Embryo
Watching and measuring what happens in tissues inside the human embryo is currently not possible, and it’s difficult to do in mammalian models. Because humans and the fruit fly Drosophila share so many biological similarities, Columbia Engineering and Syracuse University researchers tackled this problem by focusing on fruit flies. The team reports today that they can predict when the tissue will begin to rapidly flow just by looking at cell shapes in the tissue.
Researchers reveal new understandings of synthetic gene circuits
Recent discoveries by two research teams in the Ira A. Fulton Schools of Engineering at Arizona State University are advancing the field of synthetic biology. Results from a research collaboration between the lab groups of Assistant Professor Xiaojun Tian and Associate Professor have revealed novel ways that engineered gene circuits interact with biological host cells.
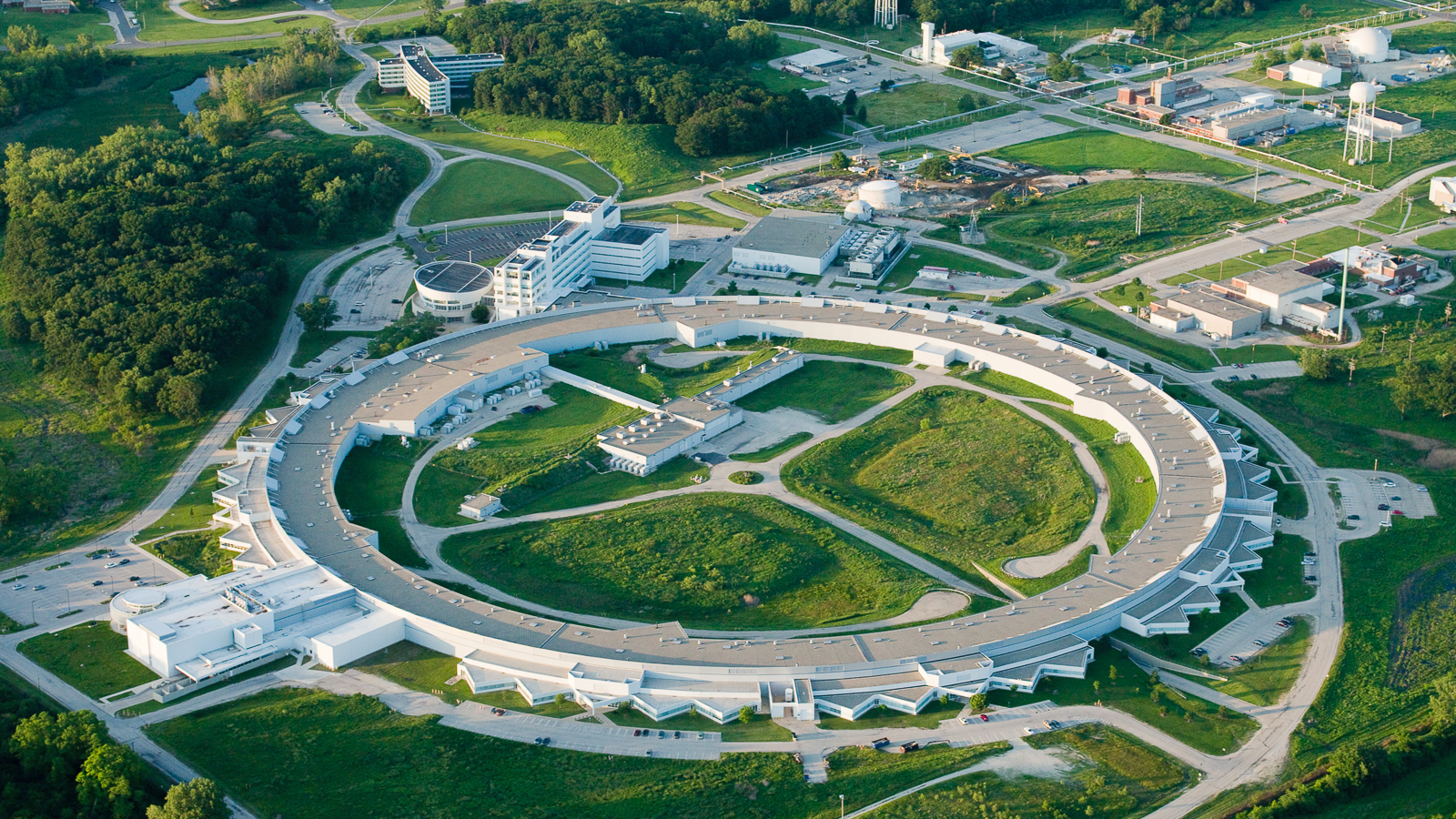

Argonne’s researchers and facilities playing a key role in the fight against COVID-19
Argonne scientists are working around the clock to analyze the virus to find new treatments and cures, predict how it will propagate through the population, and make sure that our supply chains remain intact.
PECASE Honoree Sohini Ramachandran Studies the Genetic Foundations of Traits in Diverse Populations
NIGMS grantee and presidential award recipient Sohini Ramachandran, Ph.D., is challenging our understanding of genetic variation among human populations. She discusses her research on how the genetic composition of traits and diseases varies among populations, the value of statistical and computational work in human genetics, and what this all means for patient treatment.
Sepsis: Using Big Data to Cut a Killer Down to Size
Sepsis causes nearly 270,000 deaths in the United States each year. Find out how big data approaches are helping clinicians catch it sooner, treat it better, and help survivors cope with long-term effects.

New Computational Tools Identify Alternative Splicing Changes in Aggressive Cancers
A multi-institutional group of researchers led by Children’s Hospital of Philadelphia (CHOP) has linked a strong cancer driver gene to changes in proteins that regulate alternative splicing. The researchers created new computational tools and biological model systems for the study. This collaborative research, led by Yi Xing, PhD, at CHOP and Owen Witte, MD, at the University of California, Los Angeles (UCLA), was published today in the Proceedings of the National Academy of Sciences.