The Science
Microbes are foundational for life on Earth. These tiny organisms play a major role in everything from transforming sunlight into the fundamental molecules of life. They help to produce much of the oxygen in our atmosphere. They even cycle nutrients between air and soil. Scientists are constantly finding interactions between microbes and plants, animals, and other macroscopic lifeforms. As genomic sequencing has advanced, researchers can investigate not only isolated microbes, but also whole communities of microorganisms – known as microbiomes – based on DNA found in an environment. The genomes extracted from these communities (metagenomic sequences) can identify the organisms that carry out biogeochemical processes, contribute to health or disease in human gastrointestinal microbiomes, or interact with plant roots in the rhizosphere. The Department of Energy Systems Biology Knowledgebase (KBase) recently released a suite of features and a protocol for performing sophisticated microbiome analysis that can accelerate research in microbial ecology.
The Impact
The widespread adoption of DNA sequencing in microbiology has generated huge amounts of genomic data. Researchers need computational tools to recover high-quality genomes from environmental samples to understand which organisms live in an environment and how they might interact. The combination of usability, data, and bioinformatics tools in a public online resource makes KBase a uniquely powerful web platform for performing this task. These new features in KBase will allow biologists to obtain genomes from microbiome sequences with easy-to-use software powered by Department of Energy computational resources. This will reduce the time required to process sequencing data and characterize genomes. Scientists can use KBase to collaboratively analyze genomics data and build research communities to solve common problems in microbial ecology.
Summary
Obtaining genomes of uncultivated microbes directly from the environment using DNA sequencing is a recent advance that allows scientists to discover and characterize novel organisms. Sequencing the DNA of all the microbes in a given environment produces a “metagenome.” Performing genetic analysis of metagenomes has emerged as a way to explore microbial traits and behaviors and community interactions in an environmental context. Methods for obtaining metagenome-assembled genomes (MAGs) have varying degrees of success, depending on the techniques used. An increasing number of researchers generate microbiome sequences, but many do not have ready access to the expertise, tools, and computational resources necessary to extract, evaluate, and analyze their genomes.
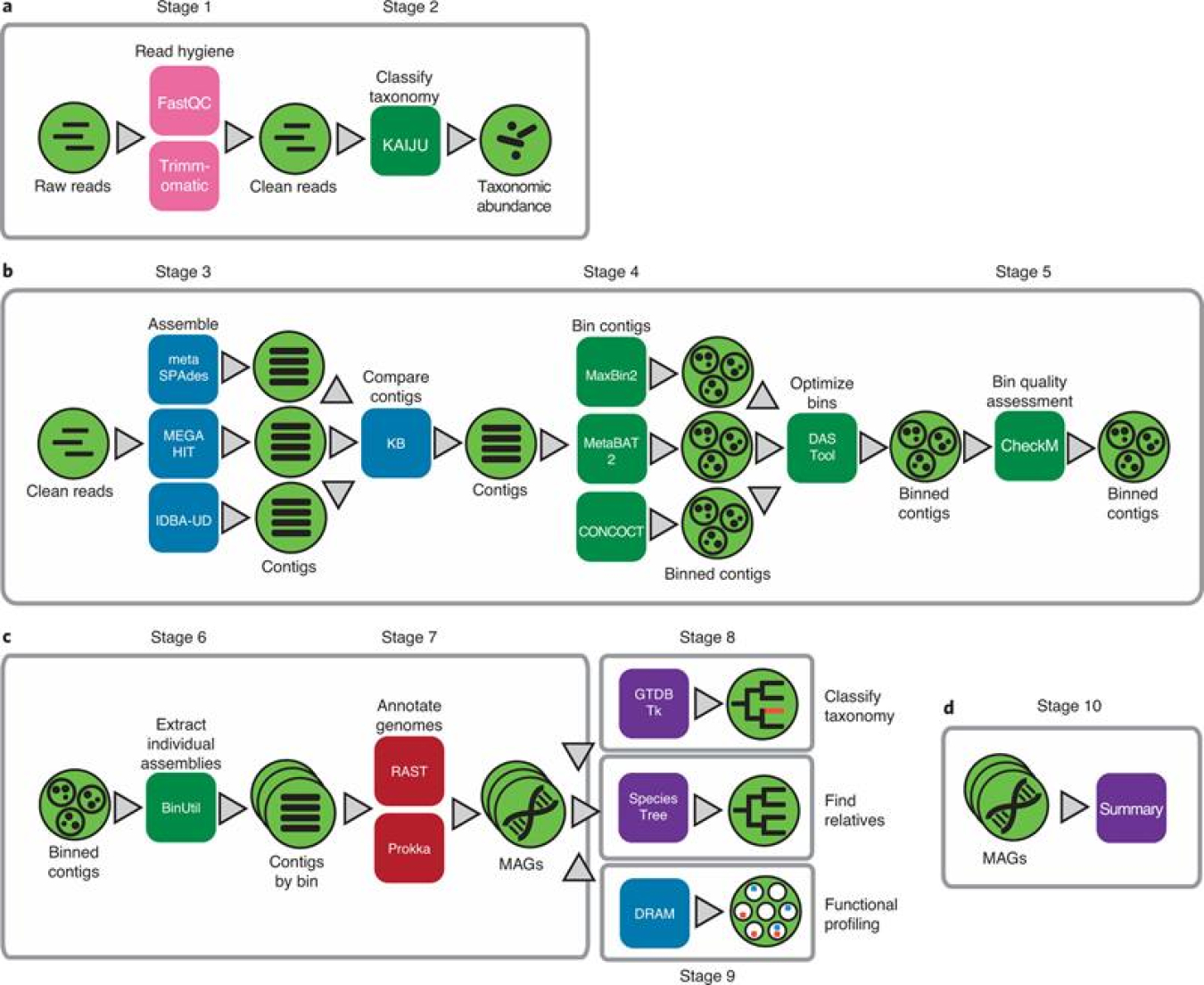
The KBase team added and updated several metagenome analysis tools, data types, and execution capabilities to provide researchers the tools that accelerate the discovery of microbial genomes and uncover the genetic potential of microbial communities. A recent paper in Nature Protocols presents a series of analysis steps, using KBase apps and data products for extracting high quality MAGs from metagenomes. These capabilities, including computing, data storage, and sharing of data and analyses, are provided free to the public via the KBase web platform. This protocol allows scientists to both generate putative genomes from organisms only found in the environment and analyze them with tools to understand who they are, what they are doing, who they are interacting with, and their role in the ecosystem.
Contact
Dylan Chivian
Lawrence Berkeley National Laboratory
[email protected]
Benjamin Allen
Oak Ridge National Laboratory
[email protected]
Funding
KBase is funded by the Genomic Science Program in the Department of Energy Office of Science, Office of Biological and Environmental Research.
Publications
Chivian, D. et al. Metagenome-assembled genome extraction and analysis from microbiomes using KBase. Nature Protocols 18 (2022). [DOI:10.1038/s41596-022-00747-x]
Related Links
Learn more about KBase: Genome Extraction from Shotgun Metagenome Sequence Data KBase Narrative
Scraped from https://www.sourcearu.com