“This is critical knowledge we need to fight future pandemics,” said Dmitry Korkin, Harold L. Jurist ’61 and Heather E. Jurist Dean’s Professor of Computer Science and lead researcher on the project. “Understanding the SARS-COV-2 virus envelope should allow us to model the actual process of the virus attaching to the cell and apply this knowledge to our understanding of the therapies at the molecular level. For instance, how can the viral activity be inhibited by antiviral drugs? How much antiviral blocking is needed to prevent virus-to-host interaction? We don’t know. But this is the best thing we can do right now—to be able to simulate actual processes.”
Feeding genetic sequencing information and massive amounts of real-world data about the pandemic virus into a supercomputer in Texas, Korkin and his team, working in partnership with a group led by Siewert-Jan Marrink at the University of Groningen, Netherlands, produced a computational model of the virus’s envelope, or outer shell, in “near atomistic detail” that had until now been beyond the reach of even the most powerful microscopes and imaging techniques.
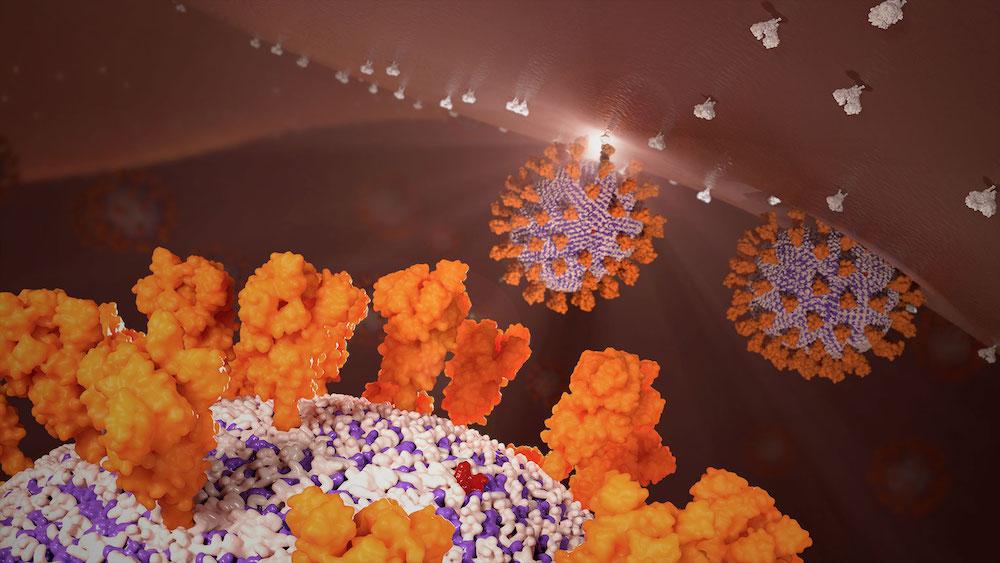
Essentially, the computer used structural bioinformatics and computational biophysics to create its own picture of what the SARS-COV-2 particle looks like. And that picture showed that the virus is more elliptical than spherical and can change its shape. Korkin said the work also led to a better understanding of the M proteins in particular: underappreciated and overlooked components of the virus’s envelope.
The M proteins form entities called dimers with a copy of each other, and play a role in the particle’s shape-shifting by keeping the structure flexible overall while providing a triangular mesh-like structure on the interior that makes it remarkably resilient, Korkin said. In contrast, on the exterior, the proteins assemble into mysterious filament-like structures that have puzzled scientists who have seen Korkin’s results, and will require further study.
Korkin said the structural model developed by the researchers expands what was already known about the envelope architecture of the SARS-COV-2 virus and previous SARS- and MERS-related outbreaks. The computational protocol used to create the model could also be applied to more rapidly model future coronaviruses, he said. A clearer picture of the virus’ structure could reveal crucial vulnerabilities.
“The envelope properties of SARS-COV-2 are likely to be similar to other coronaviruses,” he said. “Eventually, knowledge about the properties of coronavirus membrane proteins could lead to new therapies and vaccines for future viruses.”
The new findings published in Structure were three years in the making and built upon Korkin’s work in the early days of the pandemic to provide the first 3D roadmap of the virus, based on genetic sequence information from the first isolated strain in China.
About Worcester Polytechnic Institute
WPI, a global leader in project-based learning, is a distinctive, top-tier technological university founded in 1865 on the principle that students learn most effectively by applying the theory learned in the classroom to the practice of solving real-world problems. Recognized by the National Academy of Engineering with the 2016 Bernard M. Gordon Prize for Innovation in Engineering and Technology Education, WPI’s pioneering project-based curriculum engages undergraduates in solving important scientific, technological, and societal problems throughout their education and at more than 50 project centers around the world. WPI offers more than 70 bachelor’s, master’s, and doctoral degree programs across 18 academic departments in science, engineering, technology, business, the social sciences, and the humanities and arts. Its faculty and students pursue groundbreaking research to meet ongoing challenges in health and biotechnology; robotics and the internet of things; advanced materials and manufacturing; cyber, data, and security systems; learning science; and more. www.wpi.edu