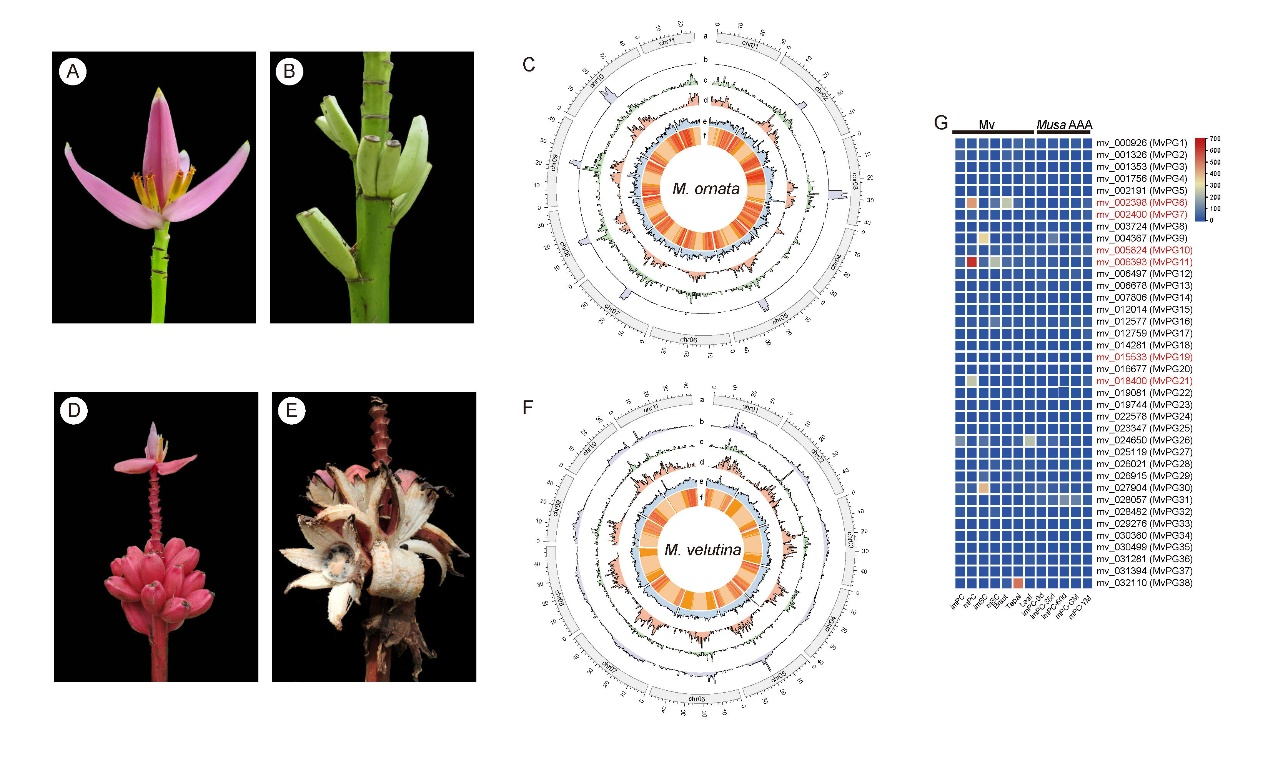
Musa ornata W. Roxburgh (Mo) and Musa velutina H. Wendl. & Drude (Mv) belong to the section Musa of the Musaceae family. They are closely related to Musa acuminata and are native to Bangladesh, Myanmar, and northeast India. These species are not only widely cultivated as popular ornamental plants in tropical regions but also contribute to the local diet through their fruit. Therefore, Mo and Mv are desirable candidates for high-quality genome sequencing to enhance future molecular breeding. In this study, the chromosomal-level genome assemblies of Mo and Mv were generated using Oxford Nanopore long reads and Hi-C reads. The genomes of Mo and Mv were assembled into 11 pseudochromosomes with genome sizes of 427.85 Mb and 478.10 Mb, respectively. Repetitive sequences comprised 46.70% and 50.91% of the total genomes of Mo and Mv, respectively. Genome quality assessments confirmed the contiguity (LAI: 13.68 and 16.81), accuracy (mapping rates of Illumina reads: 95.55% and 94.29%), and completeness (BUSCO: 98.08% and 98.51%) of the two genomes. According to the gene predictions, a total of 39,177 and 31,256 high-confidence protein-coding genes were annotated for Mo and Mv, respectively. Compared to Musa acuminata and Mv, several large inversions and translocations were observed on chr04 of Mo. Differentially expressed gene (DEG) and Gene Ontology (GO) enrichment analyses indicated that upregulated genes in the mature pericarps of Mv were mainly associated with the saccharide metabolic processes, particularly at the cell wall and extracellular region. Furthermore, several polygalacturonase (PG) genes were identified that showed higher expression levels in mature pericarps of Mv compared to other tissues, which may be accountable for pericarp dehiscence. In addition, this study also identified genes associated with anthocyanin biosynthesis pathway. Taken together, the chromosome-level genome assemblies of Mo and Mv provide valuable insights into the mechanism of pericarp dehiscence and anthocyanin biosynthesis in banana, which will significantly contribute to future genetic and molecular breeding efforts.
The article “Chromosome-level genome assemblies of Musa ornata and Musa velutina provide insights into pericarp dehiscence and anthocyanin biosynthesis in banana” has been published in Horticulture Research. This work was supported by the National Natural Science Foundation of China. For further information, please refer to: https://doi.org/10.1093/hr/uhae079.
###
References
Authors
Tian-Wen Xiao 1,5, Xin Liu 1,5,6, Ning Fu 1,5,6, Tong-Jian Liu 1,5, Zheng-Feng Wang 2,3,5, Xue-Jun Ge 4,5, Hui-Run Huang 1,5
Affiliations
1 Key Laboratory of National Forestry and Grassland Administration on Plant Conservation and Utilization in Southern China, South China Botanical Garden, Chinese Academy of Sciences, Guangzhou 510650, China
2Guangdong Provincial Key Laboratory of Applied Botany, South China Botanical Garden, Chinese Academy of Sciences, Guangzhou 510650, China
3Key Laboratory of Vegetation Restoration and Management of Degraded Ecosystems, South China Botanical Garden, Chinese Academy of Sciences, Guangzhou, 510650, China
4State Key Laboratory of Plant Diversity and Specialty Crops, South China Botanical Garden, Chinese Academy of Sciences, Guangzhou 510650, China
5South China National Botanical Garden, Guangzhou 510650, China
6University of Chinese Academy of Sciences, Beijing 100049, China