Like humans, bacteria live together in communities, sometimes lending a hand — or in the case of bacteria, a metabolite or two — to help their neighbors thrive. Understanding how bacteria interact is critical to solving growing problems such as antibiotic resistance, in which infectious bacteria form defenses to thwart the medicines used to fight them.
Now, researchers at the University of Delaware have discovered that bacteria do more than just work together. Bacterial cells from different species can combine into unique hybrid cells by fusing their cell walls and membranes and sharing cellular contents, including proteins and ribonucleic acid (RNA), the molecules which regulate gene expression and control cell metabolism. In other words, the organisms exchange material and lose part of their own identity in the process.
This unprecedented observation, which was reported on Tuesday, Sept. 1 in mBio, a journal of the American Society for Microbiology, has the potential to shed light on unexplained phenomena affecting human health, energy research, biotechnology and more.
The research team, led by Eleftherios (Terry) Papoutsakis, Unidel Eugene Du Pont Chair of Chemical and Biomolecular Engineering, studied interactions between Clostridium ljungdahlii and C. acetobutylicum. These species of bacteria work together in a syntrophic system, producing metabolites that are mutually beneficial to each other’s survival.
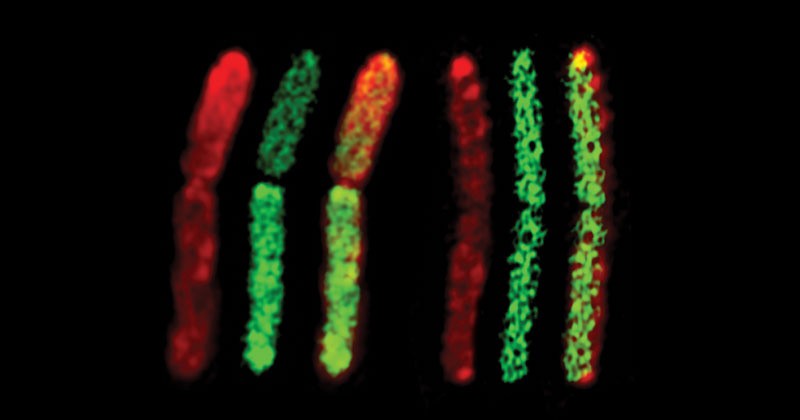
The team found that C. ljungdahlii invades C. acetobutylicum. The two organisms combine cell walls and membranes and exchange proteins and RNA to form hybrid cells, some of which continue to divide and in fact differentiate into the characteristic sporulation program.
“They mix their machinery to survive or do metabolism, and that’s kind of extraordinary, because we always assumed that each and every organism has its own independent identity and machinery,” said Papoutsakis.
Previously, researchers have observed that bacteria could exchange some material through nanotubes. The combination into hybrid cells was unexpected.
“This is the first time we’ve shown this in this bacteria, and it’s also a new mechanism of how material is exchanged,” said Kamil Charubin, a doctoral student in chemical and biomolecular Engineering and first author of the paper.
Although this phenomenon of interspecies microbial fusion is now being reported for the first time, it is likely ubiquitous in nature among many bacterial pairs.
So why do bacteria bother to fuse together? The simple answer is likely because this process allows the microbes to share machinery that will increase their odds of survival.
For example, some pathogenic bacteria — those that can cause disease — may borrow proteins from other antibiotic-resistant bacteria in order to shore up their own resistance. Some bacteria might borrow machinery from others in order to evade detection by the immune system. This could also help to explain why some bacteria are difficult to culture, or grow for study or medical diagnostic purposes. These difficult-to-culture bacteria might combine with or work with and depend on other microorganisms for their existence instead of growing and multiplying on their own.
The team’s findings may influence understanding of the evolution of biology because once bacterial species share machinery, they can evolve together instead of only evolving on their own, said Papoutsakis.
This work was supported by the Army Research Office (award no. W911NF-17-1-0343, and W911NF-19-1-0274) and the U.S. Department of Energy (DE-SC0019155).
“These findings will guide new thinking in not just the field of microbial evolution, but also toward biotechnological solutions that can benefit the soldier,” said Dr. Robert Kokoska, program manager, Army Research Office (ARO), an element of the U.S. Army Combat Capabilities Development Command’s Army Research Laboratory. “These include studies of how the human microbiome shapes soldier human health and cognition and how microbial communities can be better designed for a broad range of advances including strategies for reliable in-field biological sensing, waste remediation and novel means of biosynthesis.”
The paper’s authors also include Shannon Modla and Jeffrey Caplan of the Delaware Biotechnology Institute and the University of Delaware Bioimaging Center.