New research led by Penn State reveals that the stem region of the spike protein became progressively tighter over time, and the team thinks this likely improved the virus’s ability to transmit through nasal droplets and infect host cells once in the body. The team said the stem region of the protein that emerged in the most recent Omicron variants is as rigid as it can get, which could mean that newer vaccines may be effective for longer than the ones that targeted the original variant.
“We wanted to see how the spike protein morphed structurally as it evolved from the original wild-type strain of the virus, through the alpha, delta and most recently Omicron variants,” said Ganesh Anand, associate professor of chemistry and of biochemistry and molecular biology, Penn State. “We found that the spike protein was initially more flexible at the stem region, which is where the spike protein is bundled together, but over time, mutations caused the protein to become progressively tighter and more rigid, and we think it’s now as rigid as it can get. This is important because it means that vaccines that are developed to target the current variant with these rigid spike proteins are likely to be effective for much longer than previous vaccines against the more flexible wild-type strain.”
To study how the spike protein changed with each of the new variants, the team studied the virus in vitro (in a test tube) using a technique called amide hydrogen/deuterium exchange mass spectrometry.
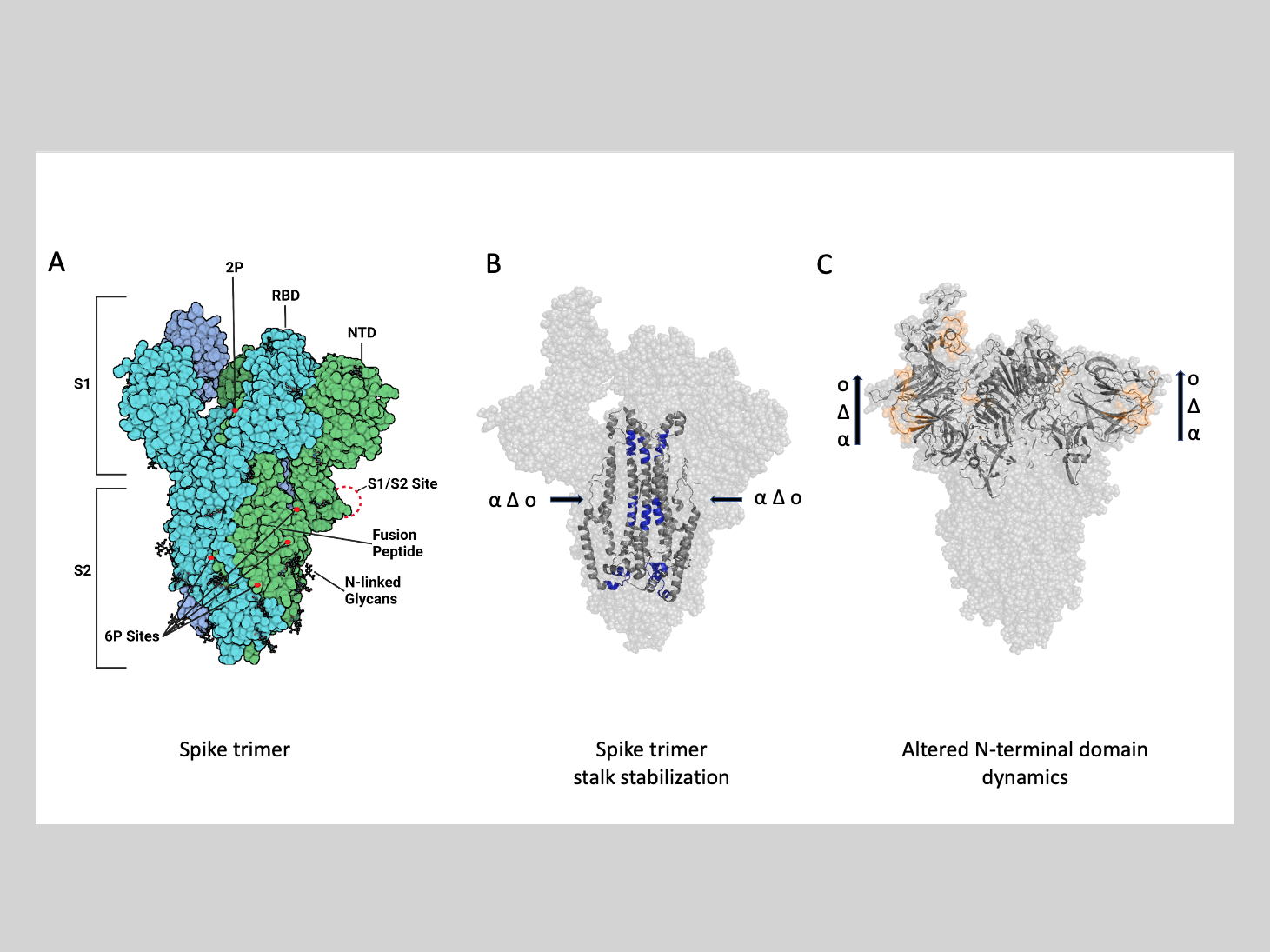
Anand explained that the SARS-CoV-2 spike protein is composed of three chain molecules called monomers that are bound together to form a trimer. The spike protein is made up of two subunits, an S1 and S2 subunit. The S1 subunit contains a receptor binding domain while the S2 subunit contains the stem region responsible for bundling the trimer.
“It is analogous to a tree, with the stem forming the trunk and the receptor binding domain forming the branches,” said Anand.
The team’s results, which published in the journal eLife, revealed that the spike protein stem first became more rigid with the D614G mutation, which is common to all SARS-CoV-2 variants. The stem became progressively more twisted with the emergence of new mutations in subsequent variants, and the Omicron BA.1 variant showed the largest magnitude increase in stabilization relative to preceding variants.
Why would the virus benefit from a tighter core?
“We did not study the virus in patients, so we cannot determine if the changes we observed in the spike protein directly affected the newer variants such as Omicron’s ability to transmit more readily; however, we can say that the changes likely made the virus more fit, which could translate to better transmission,” said Anand. “A tighter core could likely make the virus more stable in nasal droplets and faster at binding to and entering host cells. So, for example, what initially took about 11 days to develop an infection after exposure now takes only about four days.”
Anand noted that one of the reasons the vaccines have not been able to fully neutralize the virus is because they were generated against the spike protein of the original wild-type variant.
“The latest bivalent booster — which targets newer variants — helps, but people who never got this booster aren’t receiving this more targeted protection,” he said. “Future vaccines that focus specifically on Omicron are likely to be effective for longer.”
Finally, Anand said that the spike protein has now become so tightly twisted that it is unlikely to structurally change further at the stem region.
“There are limits to how much it can tighten,” he said. “I think that we can have some cautious optimism, in that we’re not going to continuously have variants emerging, at least tightening is not going to be a mechanism.”
Other Penn State authors on the paper include chemistry graduate students Sean Braet, Theresa Buckley and Varun Venkatakrishnan. Kim-Marie Dam, postdoctoral research fellow, and Pamela Bjorkman, assistant professor of biology and biological engineering, Caltech, also are authors.